cealign
A Rust implementation of the Combinatorial Extension (CE) algorithm for protein structure alignment, available as both a command-line tool and a library.
Usage
CLI
Usage: cealign [OPTIONS] -m <MOBILE> -t <TARGET>
Options:
-m <MOBILE> Path to the mobile structure
-t <TARGET> Path to the target structure
-o, --output Save aligned structures as PDB files (`<name>_aln.pdb`)
-v, --verbose Increase output verbosity
-p, --plot Save the alignment path plot as plot.png [requires --features plot]
-h, --help Print help
-V, --version Print version
Align two structures and print the RMSD:
Save the aligned structures to disk:
Save the alignment path plot (requires --features plot):
Library
use ce;
let = open.unwrap;
let = open.unwrap;
let = align;
println!;
Installation
CLI
To include the optional alignment path plot feature:
The
plotfeature requireslibfontconfigon Linux (sudo apt-get install libfontconfig-dev).
Library
Add to your Cargo.toml:
[]
= "0.1"
Example
Align crambin X-ray structure (1CRN) against the first NMR model of (1CCM).
1CCM is a multi-model NMR ensemble, so we extract model 1 first:
# Download structures
# Extract model 1 from the NMR ensemble
# Align
Expected output (RMSD in Å over the aligned Cα atoms):
1.331
Full run with verbose logging, alignment output, and plot (requires --features plot):
[DEBUG cealign] Verbose mode enabled
[INFO cealign::ce] Initial RMSD (full): 22.172
[DEBUG cealign::ce] find_path: 20 candidate paths in buffer
[DEBUG cealign::ce] Candidate path: 5 AFPs (40 residues), aligned RMSD = 1.331
...
[INFO cealign::ce] Final RMSD (aligned CA, 40 residues): 1.331
[INFO cealign::ce] Final RMSD (full): 1.921
1.331
Saved: plot.png
Saved: 1crn_aln.pdb and 1ccm_1_aln.pdb
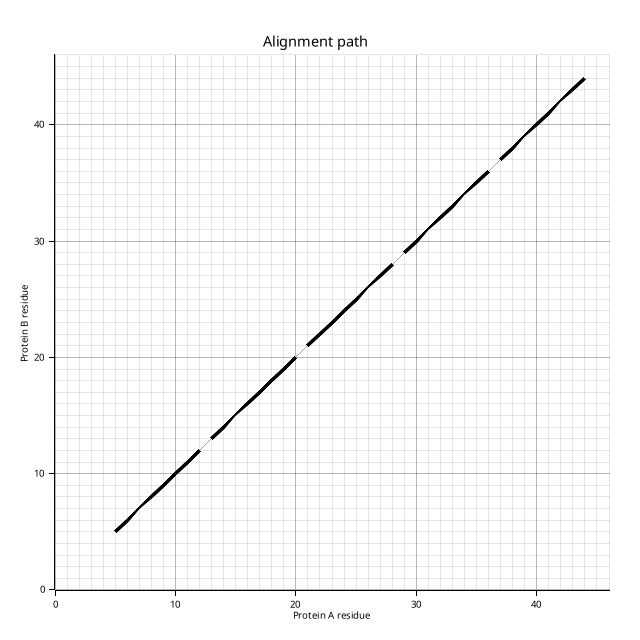
The alignment path plot (plot.png) shows each AFP as a diagonal segment on a residue-index grid, mirroring Figure 2 from Shindyalov & Bourne (1998):
Limitations
- Single-model PDB files only — multi-model NMR ensembles must be split before use
- Alignment is performed on Cα atoms; all heavy atoms are transformed using the resulting rotation and translation
See Also
- PyMOL cealign
- Biopython Bio.PDB.cealign
- Shindyalov & Bourne (1998) — original CE algorithm paper
